Abteilung Oberflächengewässer
Antibiotikaresistenzen als Umweltkontamination
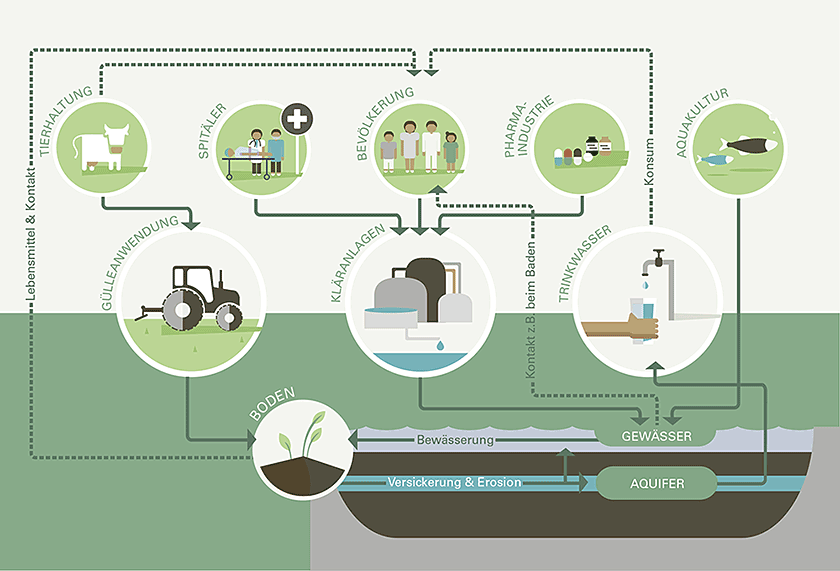
Antibiotikaresistenzen sind auf dem Vormarsch und stellen aktuell eines der größten Risiken für die öffentliche Gesundheit dar. Zunehmende Antibiotikaresistenz könnte zu einer größeren Krankheitslast, steigenden Gesundheitskosten und sogar einer höheren Sterblichkeit führen.
Umweltbakterien sind eine natürliche Quelle für Resistenzgene. Die Umwelt wird jedoch auch zunehmend durch resistente Bakterien kontaminiert, die mit behandelten oder unbehandelten Abwässern oder aus landwirtschaftlichen Quellen freigesetzt werden. Sowohl städtische als auch natürliche aquatische Systeme sind daher wichtig für das Verständnis - und die Bekämpfung - von Antibiotikaresistenzen aus einer One-Health-Perspektive.
Wir untersuchen, wie sich resistente Bakterien und ihre Resistenzgene in der aquatischen Umwelt verbreiten. Wir untersuchen, wie die Überwachung der Antibiotikaresistenz im Abwasser und in der Umwelt die Gesundheitspolitik und die Entscheidungsfindung unterstützen kann. Wir untersuchen die Dynamik der Resistenz während der Abwasserbehandlung und erforschen Strategien für eine verbesserte Eliminierung resistenter Bakterien.
Ausgewählte Publikationen
Aktuelle Projekte
Abgeschlossene Projekte
Dynamics of antibiotic resistance during oxidative treatment of wastewater
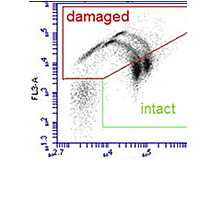
Wastewater treatments plants (WWTP) discharge antibiotic resistant bacteria with the treated wastewater. In this project we quantified the extent to which different WWTP in Switzerland eliminate and discharge resistant bacteria and how this affects the receiving waters.
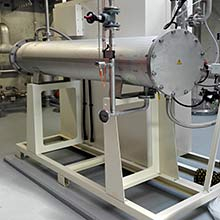
Oxidative wastewater treatment is an important strategy for the removal of micropollutants from wastewater, and is one of the main technologies that will be used for this purpose in Switzerland. In this project we studied how this treatment affects resistant bacteria. Laboratory studies were performed to obtain detailed information on how ozonation damages resistant cells and resistance genes to provide Information about dose dependence and kinetics of the process. Investigations at the first full scale ozonation plant in Switzerland, ARA Neugut, provided data from a real world implementation of this treatment process.
Key results
The project showed that ozonation of wastewater ultimately provides a less complete disinfections and a less effective deactivation of DNA (and thus resistance genes) in wastewater ozonation then simple laboratory experiments may suggest. The interactions with the complex chemistry in the wastewater matrix, microbiological populations that are less sensitive than laboratory strains such as E. coli, and the formation of larger biomass structures (flocs) all interfere. Under process conditions that are optimized for micropollutant reduction most bacterial cells are killed, but a residual population survives, and the genetic material (theoretically capable of being taken up and incorporated into the genome of other bacteria) survives. The full scale plant in Neugut revealed another potential problem – the biological post-treatment of ozonated water, which is necessary to reduce toxic oxidation products, allows resistant bacteria to grow, reducing the overall effectiveness of the process for reducing the load of resistant bacteria with the wastewater stream into rivers and lakes. Further research will be necessary to identify potential improvements to maximize the potential of wastewater ozonation as a barrier to the dissemination of antibiotic resistance into the environment.
Publications
- Czekalski et al. (2016) Antibiotikaresistenzen im Wasserkreislauf. Ein Überblick über die Situation in der Schweiz, Aqua & Gas, 96(9), 72-80. https://www.dora.lib4ri.ch/eawag/islandora/object/eawag:10653
- Czekalski et al. (2016) Inactivation of antibiotic resistant bacteria and resistance genes by ozone: from laboratory experiments to full-scale wastewater treatment, Environmental Science and Technology, 50(21), 11862-11871. https://www.dora.lib4ri.ch/eawag/islandora/object/eawag:14002
- Lee et al. (2016) Inactivation efficiency of Escherichia coli and autochthonous bacteria during ozonation of municipal wastewater effluents quantified with flow cytometry and adenosine tri-phosphate analyses, Water Research, 101, 617-627. https://www.dora.lib4ri.ch/eawag/islandora/object/eawag:10617
- Böhler et al. (2017) Biologische Nachbehandlung von kommunalem Abwasser nach Ozonung – ReTREAT. Schlussbericht, Dübendorf: Eawag. https://www.dora.lib4ri.ch/eawag/islandora/object/eawag:16638
Antibiotikaresistenz, ökotoxikologische Auswirkungen, und Risikobeurteilung von Mikroschadstoffen in einem mittelgrossen See
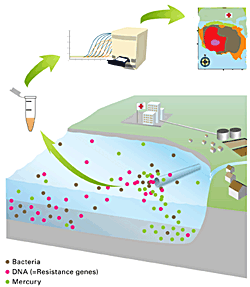
Antibiotikaresistenz wird zunehmend auch als Umweltkontamination betrachtet. In diesem Projekt wurde das Vorkommen von Antibiotikaresistenzen im Abwassersystem von Lausanne und der Einfluss der Einleitung des gereinigten Abwassers auf das Wasser und die Sedimente im Genfer See untersucht.
Zusammenarbeit
Finanzierung
Publikationen
- Bürgmann H. (2014) Eintrag von Antibiotika und Antibiotikaresistenzen in Wassersysteme der Schweiz. Prävention und Gesundheitsförderung 2014, 9:185–190, doi:10.1007/s11553-014-0444-3
- Czekalski N., Gascón Díez E., Bürgmann H. (2014) Wastewater as a point source of antibiotic-resistance genes in the sediment of a freshwater lake. The The ISME Journal (2014) 8, 1381–1390, doi:10.1038/ismej.2014.8
- Cantas L., Shah Syed Q.A., Cavaco L.M., Manaia C.M., Walsh F., Popowska M., Garelick H., Bürgmann H., Sørum H. (2013) A brief multi-disciplinary review on antimicrobial resistance in medicine and its linkage to the global environmental microbiota. Frontiers in Microbiology, May 2013, Vol. 4, Art. 96, 1-14, doi:10.3389/fmicb.2013.00096
- Czekalski N. (2013) Dissertation: Sources, Spreading and Fate of Antibiotic Resistance Genes and Resistant Bacteria in Vidy Bay, Lake Geneva, Switzerland, doi:10.5075/epfl-thesis-5637
- Czekalski N., Berthold T., Caucci S., Egli A., Bürgmann B. (2012) Increased levels of multiresistant bacteria and resistance genes after waste water treatment and their dissemination into Lake Geneva, Switzerland. Frontiers in Microbiology, March 2012, Vol. 3, Art. 106, doi:10.3389/fmicb.2012.00106
Antibiotikaresistente Mikroorganismen im Trinkwassersystem von Lausanne

In diesem Projekt wurde das Vorkommen von resistenten Bakterien im Trinkwasser von Lausanne untersucht. Über ein Jahr lang wurden Proben von Rohwasser und aufbereitetem Trinkwasser aus den verschiedenen Trinkwasserproduktionsstätten und dem Trinkwasser-Leitungs Netzwerk von Lausanne analysiert.
Finanzierung
Elimination von antibiotikaresistenten Bakterien aus Abwasser durch Membranfiltration

In diesem Projekt wurde das Vorkommen von resistenten Bakterien in der Abwasserreinigungsanalage von Lausanne und seine zeitliche Variabilität untersucht. Die effektivität der Membranfiltration bei der Elimination von resistenten Bakterien wurde quantifiziert.
Zusammenarbeit
Frederik Hammes, Umweltmikrobiologie, Eawag







