Department Environmental Chemistry
Enzyme database mining
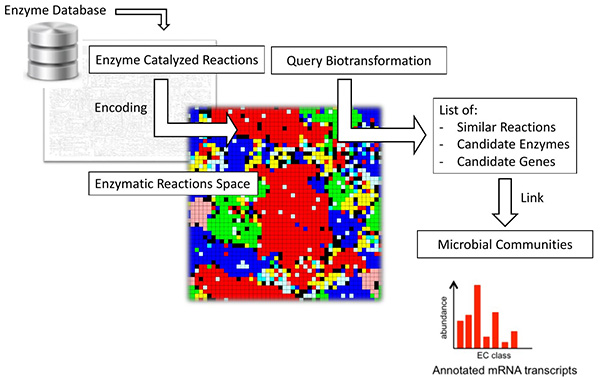
For most observed biotransformations of chemical contaminants in the environment it is not known which enzymes are catalyzing the reaction. This severely limits the possibilities to translate genomic information on microbial communities into a prediction of their ability to biotransform chemicals of interest. However, existing enzyme databases contain vast amounts of data about known enzymatic reactions that could be learned from.
In this subproject of the ERC-funded project PROduCTS , we explore the possibility to learn from data on known enzymatic reactions to predict candidate enzymes for environmental biotransformation reactions. Such an algorithm could potentially help improving the selectivity of biotransformation predictions by comparison to known substrates of similar reactions and by enabling a linkage to functional genomics-based descriptions of microbial communities.
