Department Environmental Toxicology
Toxic effects on metabolism
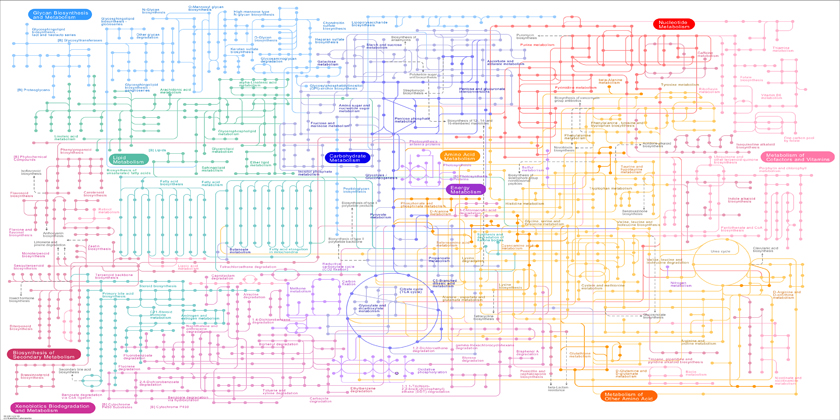
The development of mathematical models for prediction of ecological risks posed by environmental chemicals requires good understanding of toxicological mechanisms. An increasingly popular strategy for discovering new mechanisms is the use of omic technologies (toxicogenomics). However, because these approaches do not integrate mechanistic biological knowledge in the analysis, they are of limited use in moving beyond simple pathway identification. In this project, we propose a new concept for predicting the toxicity of single chemicals and chemical mixtures. This concept is based on genome-scale metabolic models (GEMs), which represent the mass-balanced entire network of metabolic reactions of a species. Based on integration with available transcriptomic and proteomic datasets, we plan to predict the effects of chemical viability, growth and energy levels of the green alga Chlamydomonas reinhardtii. We hope that this approach will enable:
- uncovering molecular mechanisms leading to adverse outcomes after exposure to single chemicals
- quantitative prediction of outcomes after exposure to single chemicals
- uncovering of molecular mechanisms leading to synergistic or antagonistic effects after exposure to chemical mixtures
- quantitative prediction of outcomes after exposure to chemical mixtures.