News Detail
Enrichment of resistance genes in wastewater treatment plants
December 12, 2018 |
Although wastewater treatment plants (WWTPs) remove over 95 per cent of human fecal bacteria, many resistant bacteria can still be detected in the final effluent. How is this to be explained? To find out, a group led by microbiologist Helmut Bürgmann investigated the fate and expression of antibacterial resistance genes in the course of treatment at twelve WWTPs. The scientists were also interested in whether the occurrence of resistance genes is influenced by stressors – such as antibiotics, biocides or heavy metals – in wastewater.
Core group of persistent resistance genes
Biomass samples were collected from the influent, the biological treatment steps and the effluent of the twelve WWTPs. DNA extracted from these samples was then analysed to identify sequences encoding antibiotic resistance. According to Bürgmann, while the levels of resistant bacteria found in treated wastewater were generally much lower than in the influent, “The relative abundance of resistance genes increases in the WWTP.”
The researchers found a wide variety of resistance genes, with the composition of the “resistome” varying widely across different WWTP compartments. However, a small group of resistance genes was found in all treatment steps. This core group traversing the WWTP is relatively abundant. Bürgmann notes that, while around 70 per cent of the resistance genes found in the influent are eliminated in the course of treatment, others arise within the WWTP: “About 40 per cent of the resistance genes in the effluent presumably originate in the activated sludge.”
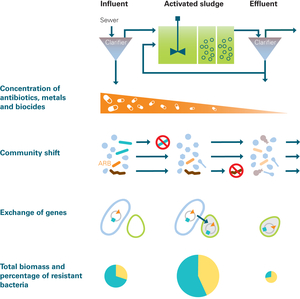
Processes possibly affecting the composition and abundance of antibiotic resistance genes during passage through a WWTP. The total amount of bacteria is highest in the biological treatment step and is much lower in the effluent than in the influent. While 95 per cent of fecal bacteria are eliminated during treatment, the proportion of resistant bacteria increases (yellow area in pie chart), i.e. resistance genes are enriched.
(Figure: Liz Ammen, Source: Bürgmann et al., 2018)
Survival through resistance
The scientists believe that environmental conditions within the WWTP offer a survival advantage for resistant microorganisms. This is suggested by the correlation observed between the abundance of resistance genes and the occurrence of certain antibiotics – even though these are only present at very low concentrations in the WWTP. In addition, resistance genes were found to be active across the entire WWTP. Bürgmann attributes the abundance of resistance genes in activated sludge bacteria to the close proximity of microorganisms during treatment: “Some bacteria in the biological treatment steps contain resistance genes with 100 per cent identity to known pathogens. These have presumably been acquired through gene exchange.”
Mobility genes
For this reason, Bürgmann’s group sought to identify not just resistance genes, but also so‑called mobility genes, which indicate that genetic material is exchanged between different bacteria. These genes were found to be frequently co-localized with resistance genes. This suggests that substantial exchanges of resistance genes occur between human pathogens and other bacteria. As Bürgmann points out, “If resistance genes are transferred to activated sludge bacteria and these enter the environment, they can presumably survive better there than the pathogens.” The easiest way of preventing this, he adds, is to eliminate biomass as far as possible from the water in the WWTP. A contribution to this goal will be made by the new treatment steps designed to remove micropollutants, which are to be introduced at Swiss WWTPs over the coming years.