Department Environmental Chemistry
PROduCTS - Predicting environment-specific biotransformation of chemical contaminants
Waste chemicals released into the environment are detrimental both to humans and the natural world. Fortunately, microbial communities in the environment can reduce the problem by breaking down the chemicals – a process called biotransformation. We know little about this process, with the result that for many chemicals released we do not know the extent to which they will be transformed.
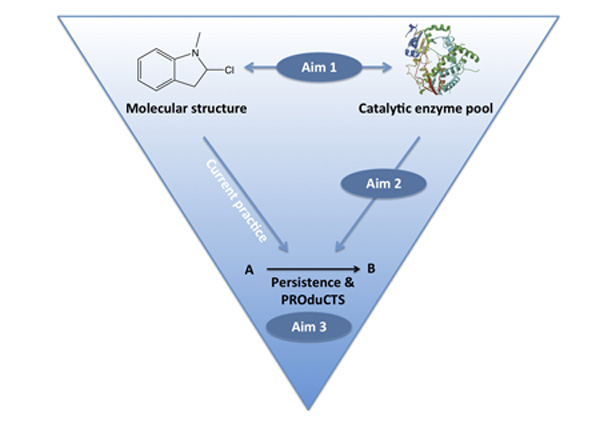
In the ERC-funded project PROduCTS we will therefore study the timescale and products of biotransformation for different chemicals, locations and microbial populations, and derive predictive methods from these observations. These will be implemented in a publically-accessible prediction system.
Our research approach is highly interdisciplinary and profits from the most recent technological and scientific advances in the fields of analytical chemistry, molecular biology and chemo-/bioinformatics. Our results should have a major impact: not only on chemical risk assessments, but also on the recovery of contaminated sites and development of green chemical alternatives.